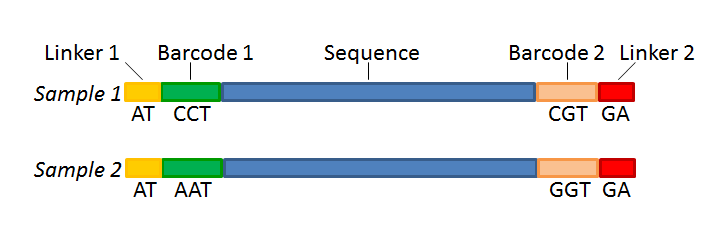
paired end sequencing reads
The standard Illumina paired-end protocol produces reads oriented pointing toward each other just like good old fashioned Sanger paired reads but the insert size is much. Biocc paired end or mate pair refers to how the library is made and then how it is sequenced.
Visit Maverix Biomics to learn more about RNA-seq.
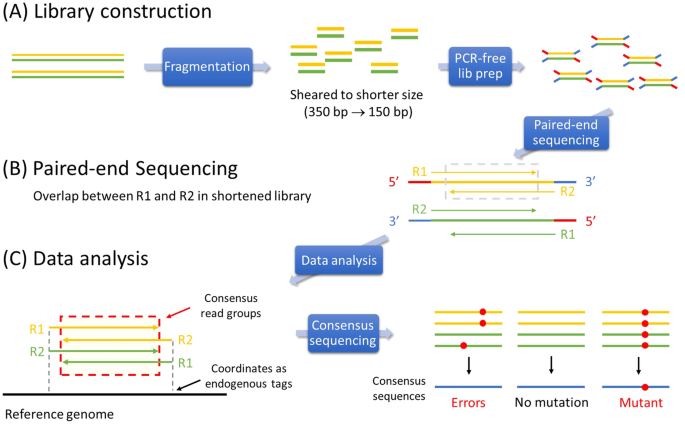
. RNA-seq analysis configuration on the Maverix Analytic Platform. Paired-end vs single-end sequencing reads. In general paired-end reads tend to be in a FR orientation have relatively small inserts 300 - 500 bp and are particularly useful for the sequencing of fragments that.
One of the advantages of paired end sequencing over single end is that it doubles the amount of data. Paired-end tags PET sometimes Paired-End diTags or simply ditags are the short sequences at the 5 and 3 ends of a DNA fragment which are unique enough that they. Paired-end DNA sequencing reads provide high-quality alignment across DNA regions containing repetitive sequences and produce long contigs for de novo sequencing by filling gaps in the.
For sequencing projects that require higher accuracy such as studies of alternate splicing 40 million to 60 million paired-end reads will provide better results. This can be done using the Set Paired Reads. Paired-end DNA sequencing reads provide high-quality alignment across DNA regions containing repetitive sequences and produce long contigs for de novo.
Paired end や mate pair という用語はどのようにライブラリが作られたかどうやってシーケンスされたかを示し. Paired end mate pair シーケンシングの解説. Adaptors P1 and P2 are ligated to both ends of the DNA molecules and the DNAs are amplified by PCR.
For more detailed analyses to. Both are methodologies that in addition to the sequence information give you. Another supposed advantage is that it leads to more accurate.
Paired-end sequencing facilitates detection of genomic rearrangements and repetitive sequence elements as well as gene fusions and novel transcripts. Read files from paired-end sequencing need to be paired in Geneious before the pairing information can be used in assembly. After the DNAs are denatured into single strands the P1 Adaptors bind to beads in the.
Paired-end 150 means that one read of 150 bases in size is generated from each end of the fragment through the inserted middle piece of target DNA from both directions for a total of 2. When you align them to the genome one read should align to the forward strand and the other should align to the reverse strand at a higher base pair position than the first one. Since paired-end reads are more likely.

Paired End Sequencing Left Showing Read 1 And Read 2 Primers Starting Download Scientific Diagram

Main Steps Of Paired End Sequencing By Illumina Technology A Download Scientific Diagram
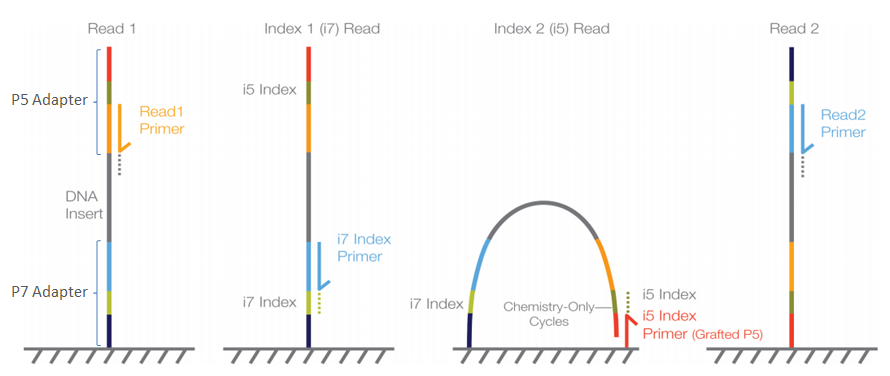
Adapter Trimming Why Are Adapter Sequences Trimmed From Only The 3a Ends Of Reads
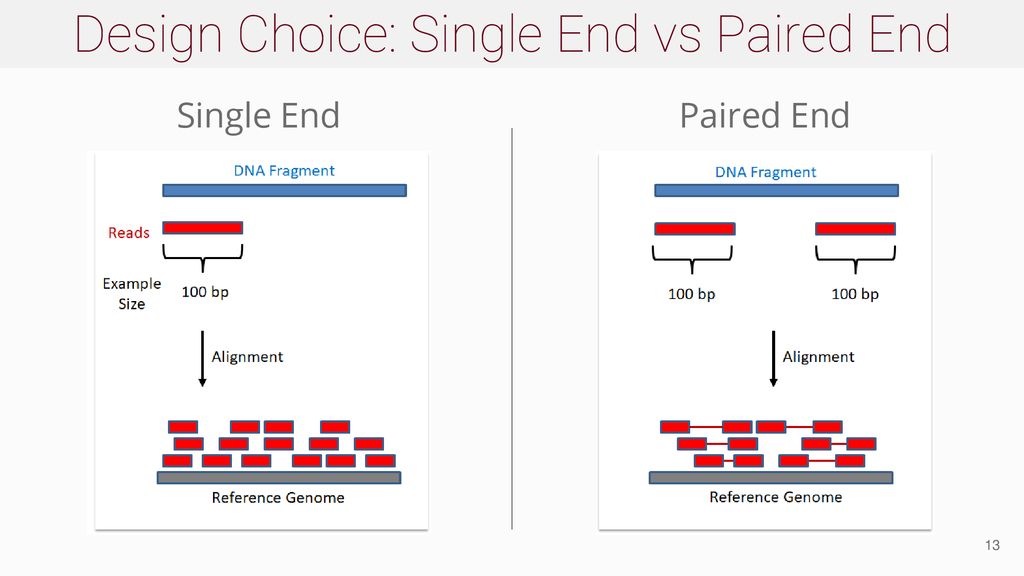
Bf Nd Next Generation Sequencing Ppt Download

4 Illumina Miseq Paired End Sequencing Steps 141 Step A To D Was Download Scientific Diagram

Long Fragments Achieve Lower Base Quality In Illumina Paired End Sequencing Biorxiv

Design Considerations Functional Genomics Ii
What Is Paired End 150 Omega Bioservices
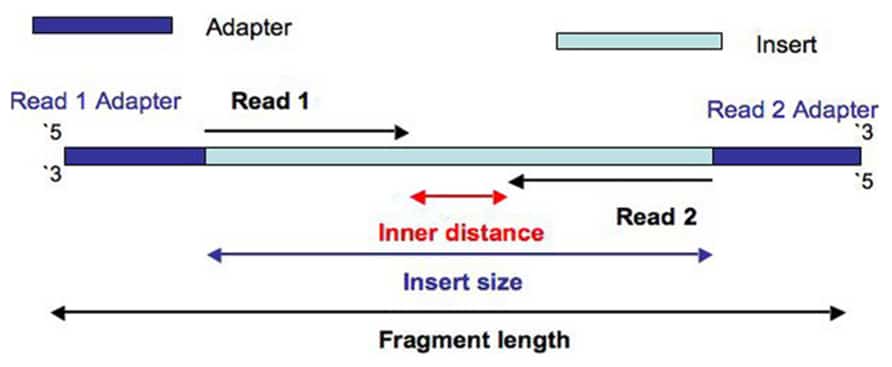
What Are Paired End Reads The Sequencing Center
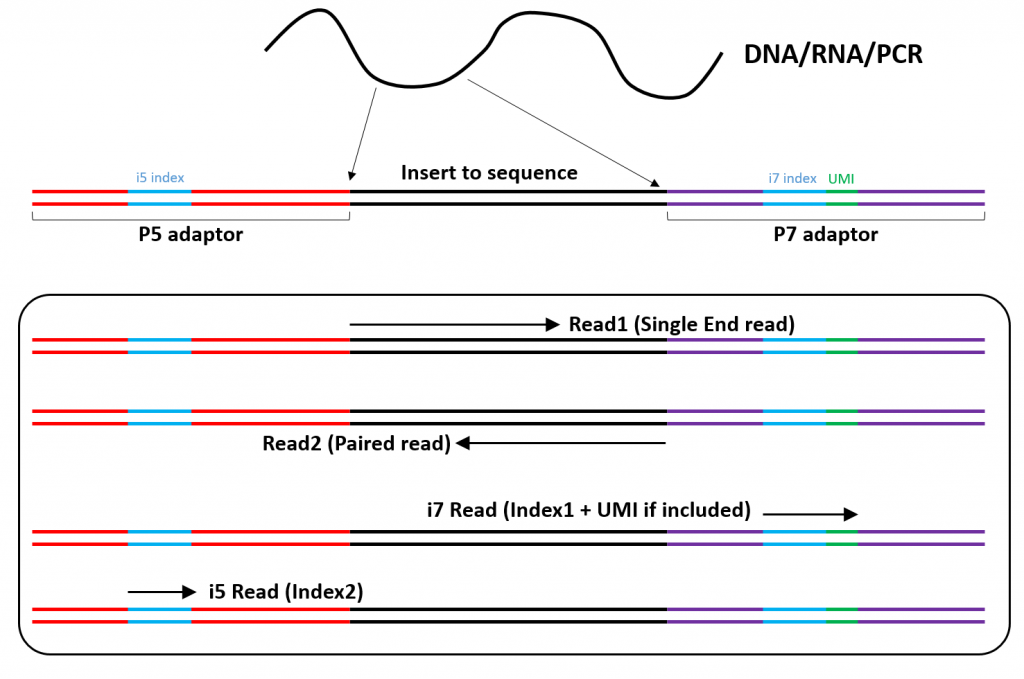
Illumina Short Read Sequencing Lausanne Genomic Technologies Facility
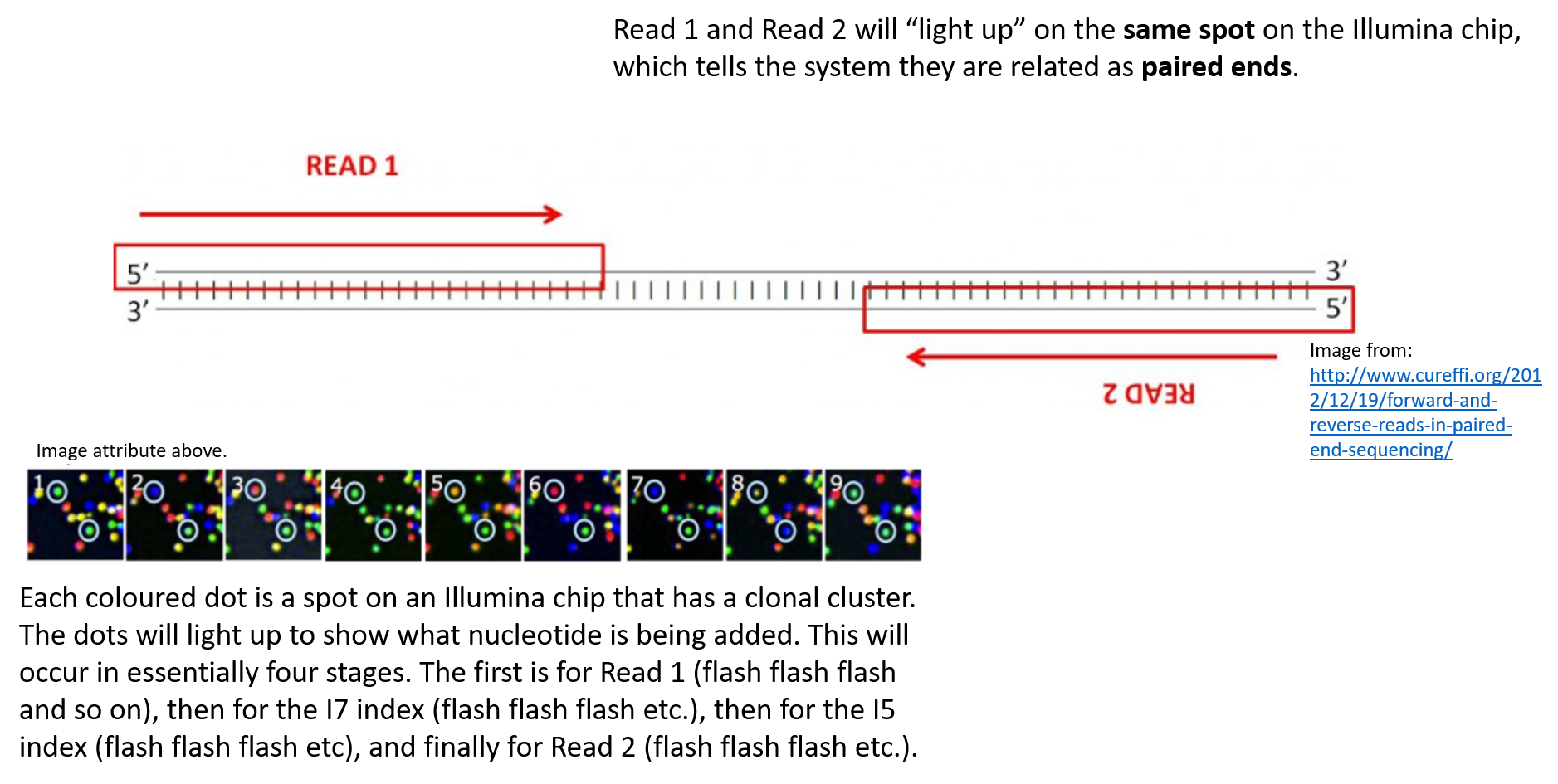
Illumina Sequencing For Dummies An Overview On How Our Samples Are Sequenced Kscbioinformatics
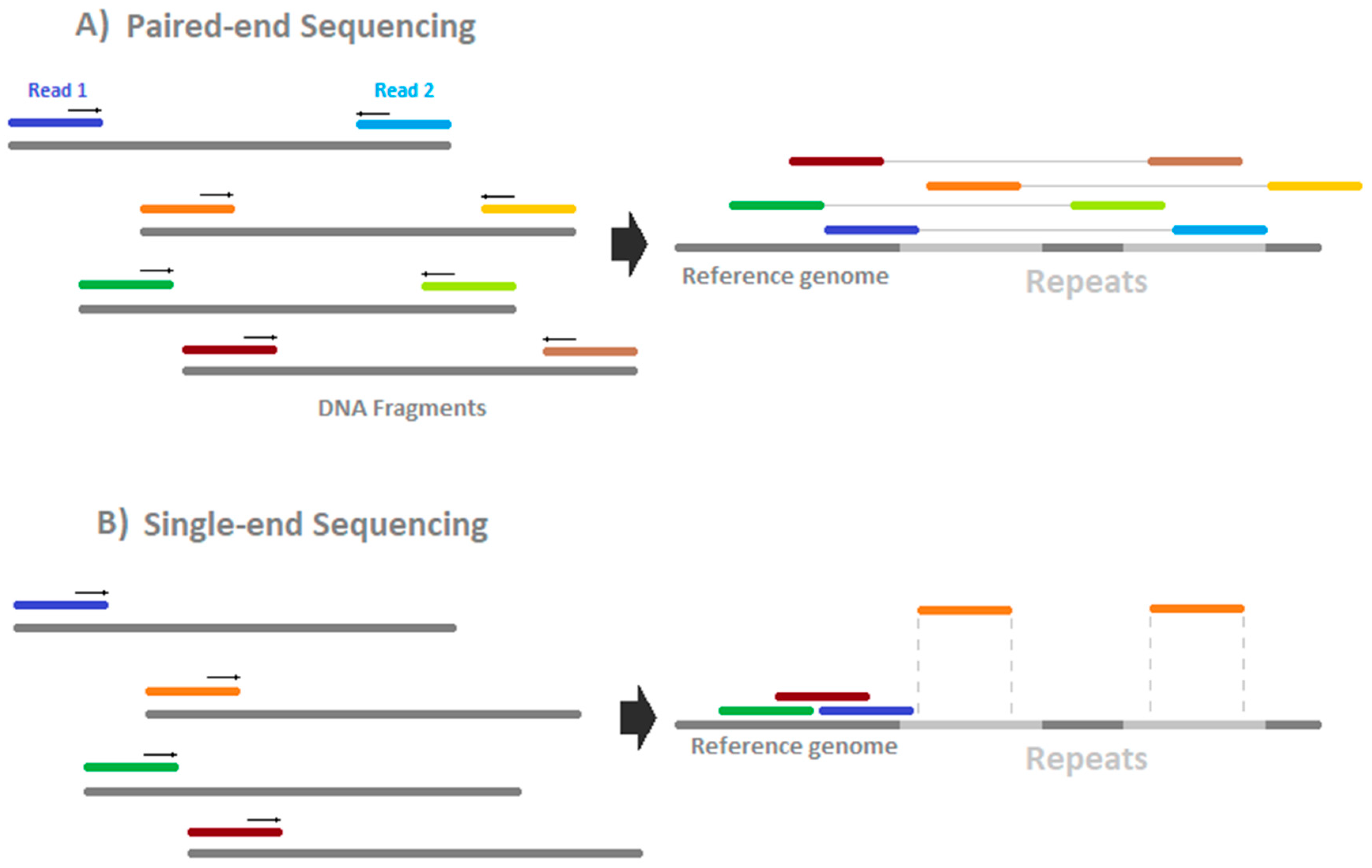
Epigenomes Free Full Text How To Design A Whole Genome Bisulfite Sequencing Experiment Html
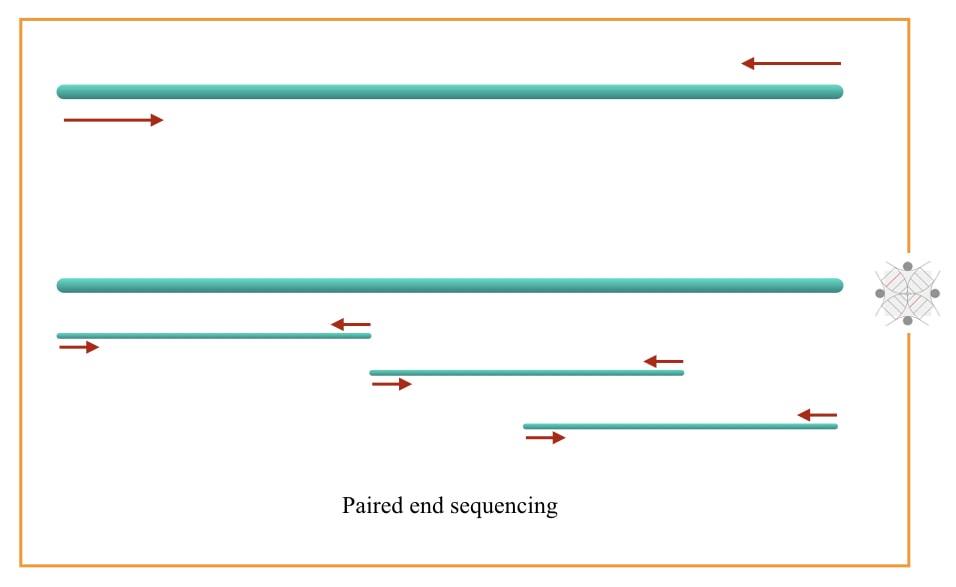
What Is Sequencing Read In Ngs Genetic Education
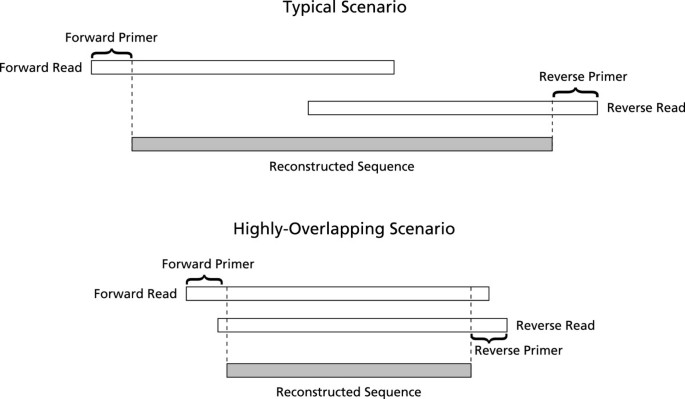
Pandaseq Paired End Assembler For Illumina Sequences Bmc Bioinformatics Full Text

Rna Seq Data Analysis 1 Raw Single End And Paired End Reads Obtained Download Scientific Diagram